Validation: Biology
Frasch (2026d). Modularity Emerges from Action-Functional Constraints in Marine Metabolic Networks — A Biology-Scale Validation of the Network-Weighted Action Principle. arXiv preprint, in submission.
The claim
If NWAP is the principle behind biological-network organisation, then natural metabolic networks at population scale should exhibit modularity excess over null models that account for upstream construction artefacts. Specifically: modularity excess $\Delta Q$ over a bipartite-incidence null — which preserves both per-protein KO count and per-KO global frequency — is the load-bearing biological observable.
Method
Tara Oceans surface and deep-chlorophyll-maximum metagenomes ($n = 7$) were processed through the MEGAHIT–Prodigal–eggNOG-mapper pipeline to recover KEGG-orthology co-occurrence networks.
Newman modularity was computed by Louvain and compared against four nulls of increasing stringency:
- Erdős–Rényi — preserves $n$ and $m$ only.
- Configuration-model — preserves the projected-graph degree sequence.
- KEGG-label permutation — preserves the per-protein KO-multiplicity distribution while shuffling KO identities.
- Bipartite-incidence null (gold standard) — randomises the underlying protein–KO bipartite graph by 2×2 edge swaps, preserving both bipartite degree sequences before re-projecting to the KO co-occurrence graph.
Recurrent multi-KO communities across the seven samples were extracted under a strict-identity definition and assigned to four pre-specified functional categories.
The result
Observed modularity is high but largely sparsity-driven ($Q \approx 0.987$, but the Erdős–Rényi null reproduces it). The biological signal is the excess over the biologically-relevant nulls:
- $\Delta Q = +0.155$ over the configuration model
- $\Delta Q = +0.179$ over KEGG-label permutation
- $\Delta Q = +0.400$ over the bipartite-incidence null ($p < 0.001$ across all seven samples)
The fine-grained communities recovered by the partition are not random: 25% recur across samples, and the 25 multi-KO communities recurring in $\geq 6/7$ samples partition cleanly into four functional categories — heterodimeric/heterocomplex enzymes, sequential biosynthetic enzymes, substrate-binding/permease pairs, and regulatory dyads / isozyme paralogs — that correspond to the cell's most biophysically- and regulatorily-constrained co-encoding units.
What this domain adds to the programme
This is the natural-system, population-scale test. It establishes that the modularity-excess signature predicted by the framework appears in field-collected metagenomic data, with the specific biological character a cost-minimisation principle predicts: assembly-stoichiometric, flux-coupled, spatially-coupled, and regulatory pairings are the co-encodings preserved across diverse marine microbial communities.
It also delivers a methodological contribution: bipartite-aware null comparison — not the absolute value of $Q$ — is the appropriate test for graph-theoretic claims of biological organisation. The absolute value is dominated by sparsity in any sufficiently fragmented network.
Code & data
- GitHub: martinfrasch/tara-modularity
- Annotations:
gs://minaction-tara-gauge-backup/annotations/latest/(public read) - Zenodo DOI: to be added at acceptance.
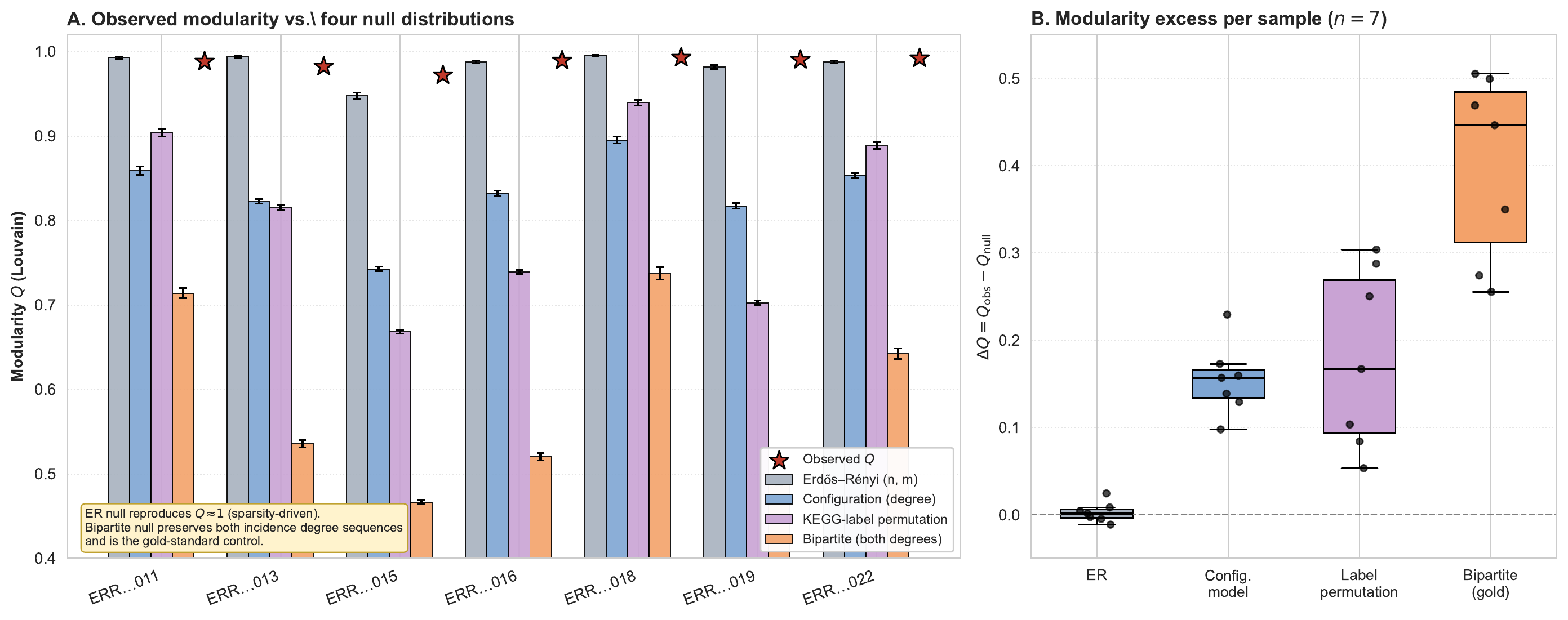
Figure 7 from the biology paper. Per-sample observed Louvain $Q$ (red star) compared with four null distributions of increasing strictness — the bipartite-incidence (gold standard) excess of $\Delta Q \approx 0.40$ is the load-bearing biological signal.